

Metagenome-Atlas¶
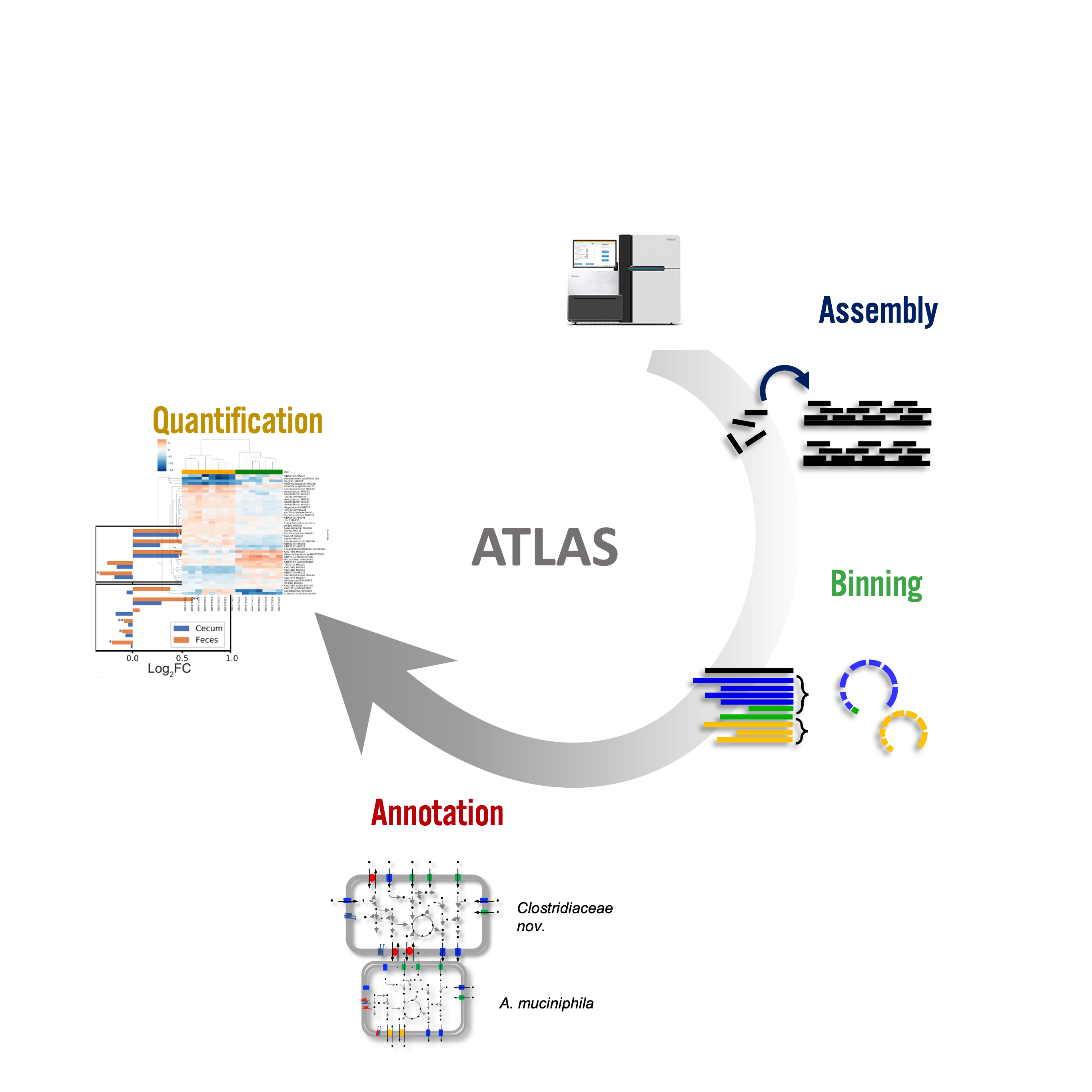
Metagenome-Atlas is a easy-to-use metagenomic pipeline based on snakemake. It handles all steps from QC, Assembly, Binning, to Annotation.
You can start using atlas with three commands:
mamba install -c bioconda -c conda-forge metagenome-atlas={latest_version}
atlas init --db-dir databases path/to/fastq/files
atlas run
where {latest_version} should be replaced by
Publication¶
ATLAS: a Snakemake workflow for assembly, annotation, and genomic binning of metagenome sequence data. Kieser, S., Brown, J., Zdobnov, E. M., Trajkovski, M. & McCue, L. A. BMC Bioinformatics 21, 257 (2020). doi: 10.1186/s12859-020-03585-4
Documentation